Adaptation and response to environmental change
~ the questions that drive me

Background
Species distributions are shaped by a combination of historical and contemporary processes, including past range expansions, migration, and patterns of adaptation to diverse environmental conditions across their ranges. Today, global climate change is a major driver of new changes in species distributions.
Warming temperatures, shifting precipitation patterns, and more frequent climate extremes have led to widespread responses like altered phenology, range shifts, and physiological changes observed across many taxa. Long-lived, sessile organisms like trees are especially vulnerable because their migration and adaptation rates cannot keep pace with rapid climate changes.
Red spruce
I used multidisciplinary approaches to understand the genotypic and phenotypic variation and climate change adaptation present in red spruce (Picea rubens Sarg.), a climate sensitive conifer species found in eastern North America. Red spruce populations have declined over the past century due to logging, acid rain, climate stress, and exhibit low genetic diversity, particularly in the fragmented southern part of its range. To assess genetic and environmental influences on key traits like phenology and growth, I planted seeds from across the species range in three common gardens (Vermont, Maryland, and North Carolina) and performed whole-genome exome capture of mother trees. Genetic analysis revealed moderate heritability and regional differentiation for phenology and growth, alongside high plasticity.
Genotypic and phenotypic variations in RS
The common gardens were raised for 2 years, recording their phenotypic and phenological traits and harvested for biomass and isotope analysis. The results of the common garden experiment to investigate the phenotypic plasticity and genotypic variations in red spruce was published in 2022. The results of this paper laid the foundations for extending our understanding of the species and further apply it in the field.
Background
We often think of species as being reproductively isolated from one another, this is not always the case. As a consequence of weak reproductive barriers, members of different species can interbreed in nature. When hybrids between two species are fertile, hybrids may then interbreed with members of the original species in a process known as backcrossing. Multiple generations of hybridization and subsequent backcrossing can facilitate the exchange of genetic material between parental species, a process known as introgression.
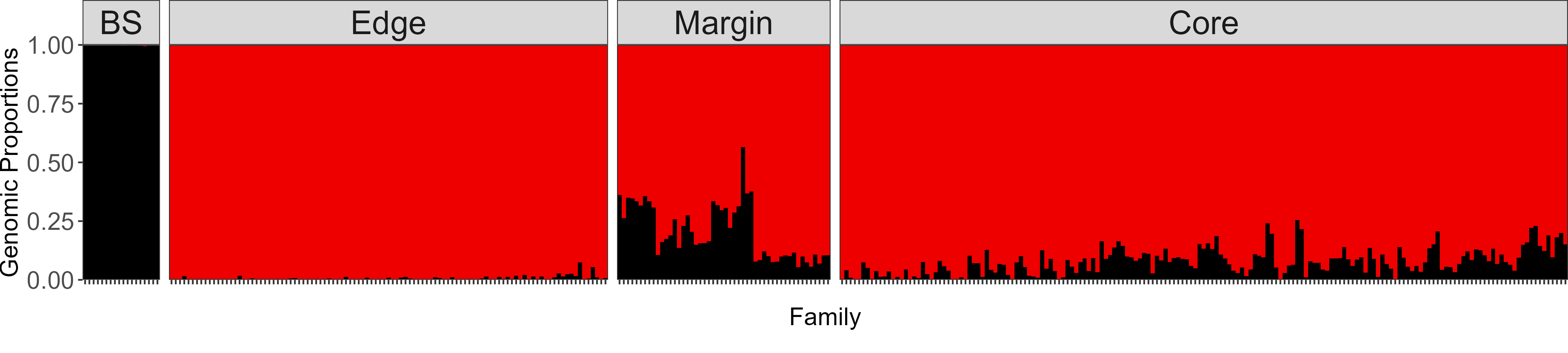
Introgression in red spruce
Northern red spruce populations shows extensive introgression from black spruce (Picea mariana), which contributes adaptive genetic diversity for key traits such as biomass and height. These advanced-generation hybrids may play a key role in climate adaptation and evolutionary rescue for red spruce populations in a changing environment. This work advances our understanding of climate adaptation in red spruce and highlights the importance of introgression as a source of adaptive genetic diversity for vulnerable forest tree species.
Background
Long‑lived, sessile organisms like forest trees have long generation times and limited dispersal, so they may not keep pace with rapid anthropogenic climate change, making climate maladaptation a major concern over the coming decades. Genomic offset provides a way to integrate genomic data and climate projections to forecast where adaptive mismatches are likely to emerge, helping identify vulnerable populations, evaluate seed transfer and assisted migration strategies, and prioritize conservation and restoration actions under changing climates.
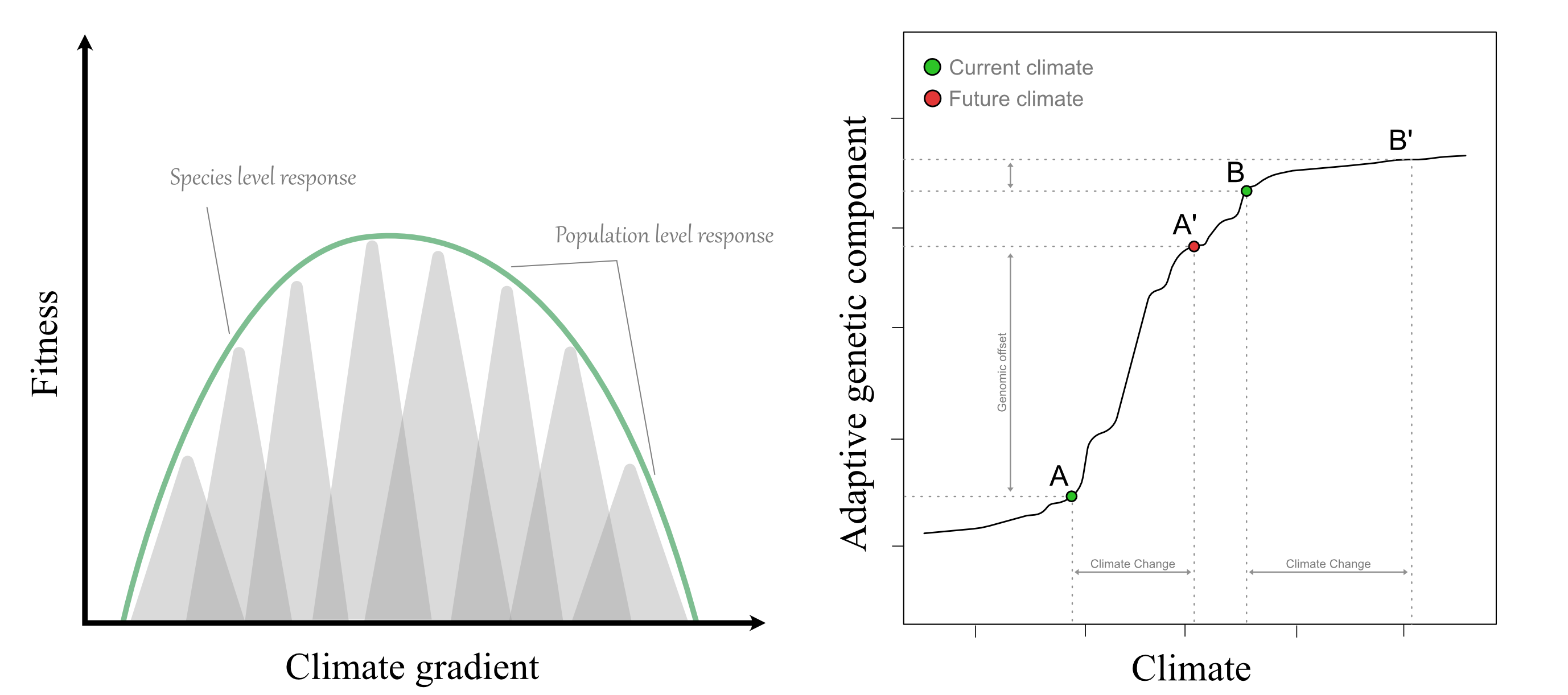
What is genomic offset?
Genomic offset is a quantitative measure of how much the allele frequencies underlying local climate adaptation would need to change for a population to remain adapted under a new climate. It is typically estimated by fitting genotype–environment association or genomic prediction models under current conditions, then projecting those models onto future or novel climates to compute the expected shift in climate‑adaptive genomic composition at each location. Large genomic offset values indicate a strong mismatch between the current genomic makeup and the climate it will experience, implying high risk of climate maladaptation.
Red spruce restoration in collaboration with CASRI and TNC
Kathryn Shallows from TNC approached the Keller lab in 2019 regarding red spruce restoration towards the southern range edge of its distribution, mainly in Maryland, Virginia and West Virginia. This led to a larger partnership that culminated in genomic assisted seed source selection for the restoration initiative at these sites. Instead of opting for a single source restoration initiative, we used a multiple seed source approach to improve the genetic diversity of the restoration stands to withstand the changing climate and environmental stressors. This source selection was carried out in the light of research done on red spruce using the exome capture data and the common gardens set up at Vermont, Maryland and North Carolina. Three to four sources combinations per restoration site were selected for high genetic diversity and low genetic load.
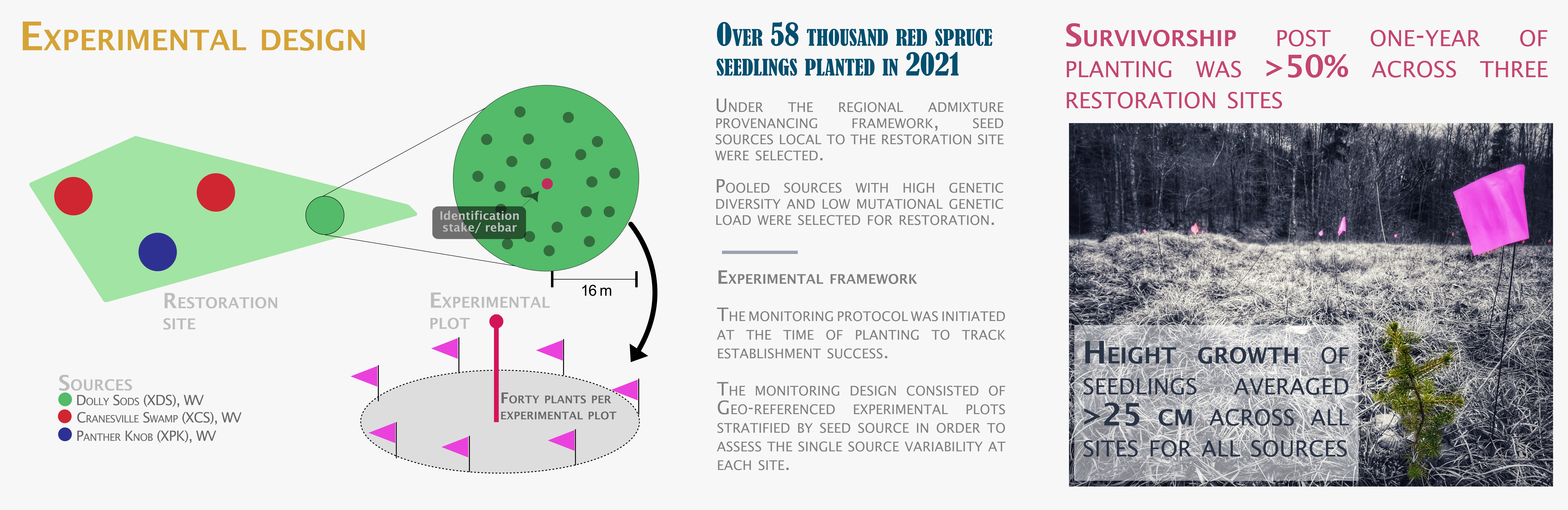
Seed sources: No one for all
Capblancq et al. (2021) has observed that early-life fitness of the red spruce had strong positive association with genetic diversity and negative association with genetic load, especially for the southern range edge of the red spruce distribution range. These seed source combinations were used as a recommendation list to Dave Seville (CASRI) to assess the seed production situation on the ground and raise them in the nursery for the restoration initiative. In 2021, TNC planted 58,000 red spruce seedlings at the restoration sites. Keller lab then carried out the restoration monitoring of these restoration sites in 2022, a whole year after the seedlings had been planted in the ground to assess the success of genomic assisted restoration of red spruce. Using this information, Prakash et al. (2024) found that there were no “Super Seed Source” that outperformed other seed sources and the combination of seed sources outperformed any single source for the restoration success.